Probabilistic data structures. Bloom filter
In the article we consider such popular implementation of a probabilistic set as Bloom filter, that can efficiently solve the problem of determining membership of some element in a large set of elements without the need to store every element and use many comparisons.
The problem
Many everyday applications require determining membership of an element in a large set of elements. Consider a problem when you develop an extension to protect users of your network from being hacked because of visiting a suspicious website. In this case, you manage a list of malware websites' URLs and every time user enters a new URL you need efficiently check it and decide whenever you allow or deny access to it.
Intuition
The simplest solution is to store all bad URLs and then search for the entered URL throw the list using your favourite search algorithm. Of course, it could have many optimization directions, but the main problem that you have to "store" them and the storage space will grow linearly within the number of elements you have. Also, search algorithms depend on the number of elements and take in average `O(log n)` comparisons (on pre-sorted data).
But wait, do you really need to know exactly which URL from your list has been entered by the user or just want to know whenever it's good or bad? It's enough to have a data structure where we don't need to store every element we know, but can fast determine whenever a new element in / not in the set using a constant time and that is not dependent on the number of documents in the set.
The basic idea is to use a limited number of fingerprints for URLs and store only them. If fingerprints are small (smaller than the URL and, usually, just an integer), this will reduce the amount of space we need to store the set, but still, it will grow linearly ... To have this space also limited we need to find such functions for fingerprints that have limited range of output values (e.g. converts every URL into an integer in the interval `[0, 2^10)`).
Cool, but everything has its own price - if we limit the number of values in output (use shorted fingerprints), we likely to have a collision (when different elements mapped to the same fingerprint), because there are much more URLs than, for instance, `2^10`, so at some point we can find 2 different URLs that are mapped to the same single value. This is why we need to use more than 1 function at the same time, it will decrease the probability of such collision significantly, but it will require more space and number of comparisons. In the end, we just need to find our "golden mean" when the probability of the collision is less important than the space/speed issues.
As soon as a new URL comes, you just need to compare the values of its fingerprints (which is limited number) and if all of them are in the known list, that you can say that the URL likely in the list, so is a bad URL.
Above I just described the idea of the most popular probabilistic data structure - a Bloom Filter, but now let's go in more details.
Bloom Filter
Bloom filter has been proposed by Burton Howard Bloom in 1970. It is a space-efficient probabilistic data structure that implements a probabilistic set with 3 operations:
- add element into the set
- test whether an element is a member of the set
- test whether an element is not a member of the set
Bloom filter can be described by 2 parameters:
- `m` - length of the filter(it is proportional to the expected number of elements `n`)
- `k` - number of different hash functions (it is assumed that `k` is much smaller than `m`)
Algorithm
Bloom filter is represented by a bit array of `m` bits. At the beginning all bits set to 0.
To insert element into the filter, we calculate values of all `k` hash functions for the element and use their output values as indices in the array where we set bits to 1.
To test if element is in the filter, we calculate all `k` hash functions for the element and check bits in all corresponding indices:
- if all bits are set, then answer is "maybe"
- if at least 1 bit isn't set, then answer is "definitely not"
Time needed to insert or test elements is a fixed constant `O(k)`, independent from the number of items already in the filter.
Bloom filter doesn't store elements themselves (only their "fingerprints" from the hash functions) and require about 1 byte per stored data.
Example
Consider Bloom filter of 16 bits (m=16):
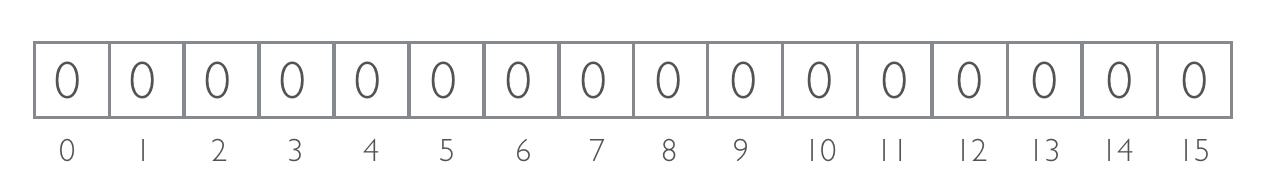
As an example, consider 2 hash functions (`k = 2`):MurmurHash3 and Fowler-Noll-Vo (to calculate the appropriate index, we divide result by mod 16)
Now, let's add element ferret to the filter: `"MurmurHash3(ferret)" = 1`, `"FNV(ferret)" = 11`. Therefore, we need to set bits #1 and #11 in the filter:
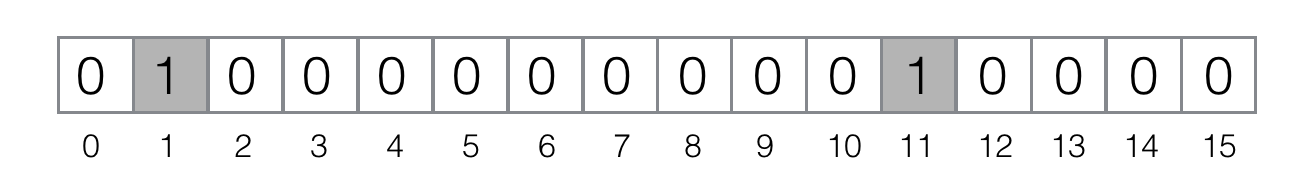
Add element bernau to the filter: `"MurmurHash3(bernau)" = 4`, `"FNV(bernau)" = 4`. This is an example of a hash collision on the functions level (it's likely, because we project all possible values the hash functions can return to the really small segment `[0, 15]`). So, we need to set only the bit #4:
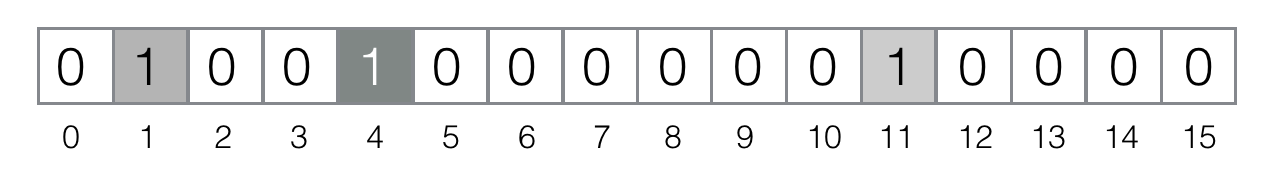
Now, let's test element berlin: `"MurmurHash3(berlin)" = 4`, `"FNV(berlin)" = 12`.Because the bit #12 is not set in the filter, we can conclude that berlin is definitely not in the set.
But that about element paris? Its hashes are `"MurmurHash3(paris)" = 11`, `"FNV(paris)" = 4`.If we check bits #4 and #11, they all set in the filter and we shell conclude that paris maybe in the set. This is an example of a false positive response, because we know that we didn't add paris to the filter and the appropriate bits were set independently by 2 different elements (bit #4 were set by bernau and bit #12 by ferret). Of course, the bigger the filter and number of hash functions, the harder to show such collision because its probability decreases.
Properties
-
False positives are possible. The situations as we saw above, when an element is not a
member, but filter returns like it is a member. Fortunately, it's not too probable situation and
we
can even estimate it's probability:
`P(e \in S| e \notin S) \approx \(1 - e^\frac{kn}{m}\)^k`
- It's possible to tune probability of false positives responses. As we can see from the formula above, such probability depends on the choice of `k` and `m`.
-
False negatives are not possible.Filter returns that an element isn't a member only if
it's
definitelly not a member:
`P(e \notin S| e \in S) = 0`
- Hash functions should be independent and uniformly distributed. In this way we randomize the hash values (set bits) uniformly in the filter and decrease probability of hash collisions on the functions level.
- Hash functions should be as fast as possible. We need to calculate all `k` functions each time we want to add a new element or test an element, therefor the faster the functions calculation, the faster our data structure can behave. The outcome: don't use cryptographic hash functions, e.g. sha1 and others.
Application
Google BigTable, Apache HBase and Apache Cassandra use Bloom filters to reduce the disk lookups for non-existent rows or columns.
Medium uses Bloom filters to avoid recommending articles a user has previously read
Google Chrome web browser used to use a Bloom filter to identify malicious URLs (moved to PrefixSet, Issue #71832)
The Squid Web Proxy Cache uses Bloom filters for cache digests
Problems
- The basic Bloom filter doesn't support deletion.
-
Bloom filters work well when they fit in main memory
- ~1 byte per element (3x-4x times more for Counting Bloom filters, that support deletion)
-
What goes wrong when Bloom filters grow too big to fit in main memory?
- On disks with rotating platters and moving heads, Bloom filters choke. A rotational disk performs only 100–200 (random) I/Os per second, and each Bloom filter operation requires multiple I/Os.
- On flash-based solid-state drives, Bloom filters achieve only hundreds of operations per second in contrast to the order of a million per second in main memory.
- Buffering can help. However, buffering scales poorly as the Bloom-filter size increases compared to the in-memory buffer size, resulting in only a few buffered updates per flash page on average.
Variants
- Attenuated Bloom filters use arrays of Bloom filters to store shortest path distance information
- Spectral Bloom filters extend the data structure to support estimates of frequencies
- Counting Bloom filters contain in each entry in the filter small counter instead of a single bit. This modification can easily support insertions and deletions by increments/decrements of the counters.
- Compressed Bloom filters can be easily adjusted to the desired tradeoff between size and false-positive rate
- Scalable Bloom filters can adapt dynamically to the number of elements stored, while assuring a maximum false-positive probability
- Bloom Filter Cascade implements filtering by cascading pipeline of Bloom filters
Open-source implementations in Python
- pybloom - A module that includes a Bloom Filter data structure along with an implementation of Scalable Bloom Filters.
- pyreBloom - Provides Redis backed Bloom Filter using GETBIT and SETBIT.
Read More
- Burton H. Bloom Space/Time Trade-offs in Hash Coding with Allowable Errors. In Communications of the ACM (1970). Vol. 13 (7). Pp. 422-426
- Wikipedia page about Bloom Filter.
- Paulo Almeida et al. Scalable Bloom Filters. In Information Processing Letters (2007). Vol. 101 (6). Pp. 255-261
- Andrei Broder, Michael Mitzenmacher Network Applications of Bloom Filters: A Survey. In Internet Mathematics (2004). Vol. 1 (4). Pp. 485-509
